UNL Bayer Hackathon - October 24-26, 2025
How to share this page
For easier sharing, the short link for this page is: https://go.unl.edu/hcc_bayer_hack_info_page

Before the Hackathon and as you arrive:¶
Summary of steps¶
- ✅ Can sign into Swan OOD
- ✅ DUO authentication tested
- ✅ Able to launch Jupyter or R Studio
- ✅ Joined Hackathon Discord
Make sure you can sign into Swan¶
Warning
Please make sure you can sign into Swan using the steps below!
The Holland Computing Center (HCC) will have very limited support over the weekend for emergencies only and may not process account requests or DUO requests during the weekend.
If you can complete the steps below, you are ready to continue.
Please make sure to complete the setup steps below.
-
Ensure your HCC Account is active.
-
Sign into Swan's Open OnDemand Portal: https://swan-ood.unl.edu
-
When you have signed into the Swan Open OnDemand Portal, please put up your Yellow Sticky Note or Green Checkmark.
If you have any issues logging in or accesing Swan, please visit our event troubleshooting guide here
Access the Hackathon Swan reservations¶
Accessing reserved Swan resources
Resources on Swan have been reserved for the period of the Hackathon. CPU-based resources consisting of 56 cores and 250GB RAM has been reserved for each team. A shared pool of GPU resources consisting of 6 NVIDIA T4 and 4 NVIDIA L40S cards has also been reserved. These GPU resources are available to all teams on a first come, first served basis.
CPU resources¶
Partition Selection
To use your team's CPU resources, you will need to ensure that your partition is set as batch
QoS
If you are getting an error when submitting a Jupyter Session or R Studio session, please make sure that your QoS Type is set to normal.
To access the reserved CPU resources, use the --reservation argument with your team number. In your SLURM script,
add the line
#SBATCH --reservation=teamX
X with your team number. For example for Team 1
#SBATCH --reservation=team1
GPU resources¶
To access the shared pool of GPU resources use the following lines
#SBATCH --reservation=gpuhack
#SBATCH --gres=gpu
Access from Open OnDemand¶
You may also access these reserved resources via Swan's Open OnDemand portal; for example to run a Jupyter notebook.
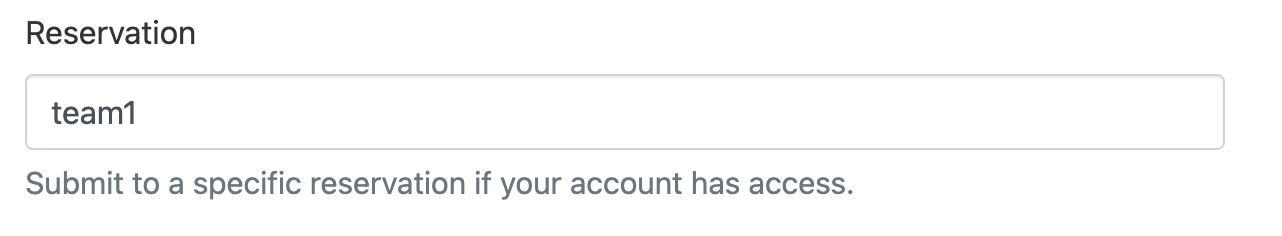
The submit form has Reservation and GRES fields which may be used identically to the --reservation argument.
For example to request the CPU resources for Team 1:
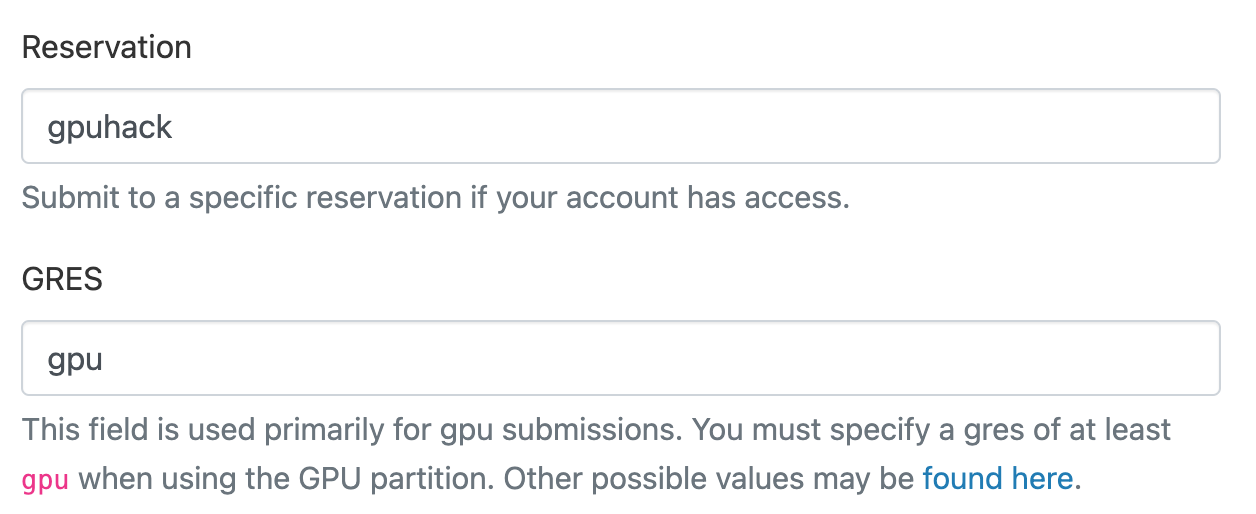
And to request the shared GPU resources
Access the team shared folder and Hackathon dataset¶
Shared folder per team¶
A shared folder per team has been created to make sharing work easier. Files created in this folder will be visible
to the entire team. Each team may only access their own folder (i.e. Team 1 cannot see Team 2's folder, etc.)
The folders are located at /work/unlbayerhack/shared/teamX where X is your team number. For example, Team 1's
folder is at /work/unlbayerhack/shared/team1. These folders may be accessed both from the command line and
from Open OnDemand (JupyterLab, RStudio, etc.).
Data Permissions
The /work/unlbayerhack/shared/teamX/Simplified Hackathon Dataset V3 directory is read-only. To work with the data, please copy it elsewhere.
Prestaged dataset¶
A copy of the Hackathon dataset has been prestaged into this folder. This copy may not be changed, but participants may make any other copies as desired.
Make sure you can launch a Jupyter Lab Session¶
JupyterLab is available for use within Open OnDemand as an Interactive App.
Please make sure you are able to launch a Jupyter session.
Configuration Details are available in the Open OnDemand Apps section
Join the Discord¶
Need help during the hackathon?
If you encounter issues with logging in, software, or jobs, please feel free to ask an HCC staff member in the room or ask in the Discord.
HCC staff will only be available on-site Friday 6pm-8:30pm and Saturday 9am - 6pm. Outside of those hours, HCC staff will be unavailable except for emergencies — see the warning above for details.
General Information¶
Various faculty members from the university and some HCC staff will be present throughout the hackathon. To help aid in communication, Sticky Notes will be used on laptops to help signify if help is needed or if everything is good.
Sticky Notes and Reactions¶
Throughout the workshop, we will use sticky notes to indicate status. These help instructors and helpers in the workshop assist with various issues and provide formative feedback during the workshops.
Please make sure the sticky note is visibly on both the front and back of your laptop
- Yellow Sticky Note: All good to go
- Red Sticky Note: I need help or I am not ready to continue
Please ask if you need help
If you are stuck or not sure on something at any point of the workshop, please let us know and we would be happy to help.
Join the Hackathon Discord!
For communication, we will be using Discord.
Useful Information and Links¶
Swan Supercomputer Login: https://swan-ood.unl.edu
Cheatsheets¶
- HCC and Slurm Cheatsheets: Sharepoint link
- Presentation Template: Link
- Bash: https://cheatography.com/davechild/cheat-sheets/linux-command-line/
- Git: https://education.github.com/git-cheat-sheet-education.pdf
- Anaconda: https://docs.conda.io/projects/conda/en/4.6.0/_downloads/52a95608c49671267e40c689e0bc00ca/conda-cheatsheet.pdf
- Python: https://indico.cern.ch/event/865287/attachments/1971788/3280306/beginners_python_cheat_sheet_pcc_all.pdf
Submitting Jobs¶
What is a supercomputer?
A supercomputer or cluster, such as Swan, comprises of many components that can be broken down into 4 major categories:
- Login Node: Where you log in to interact with the cluster. Meant for small tasks like writing files and submiting workflows.
- Head/Management Node: Manages the accounting, authentication, and management of workflows and nodes in the cluster.
- Worker Nodes: The actual powerhouse behind any cluster and run the workflows submitted from the login node.
- Filesystems: One or more storage locations for data, scripts, and output results.
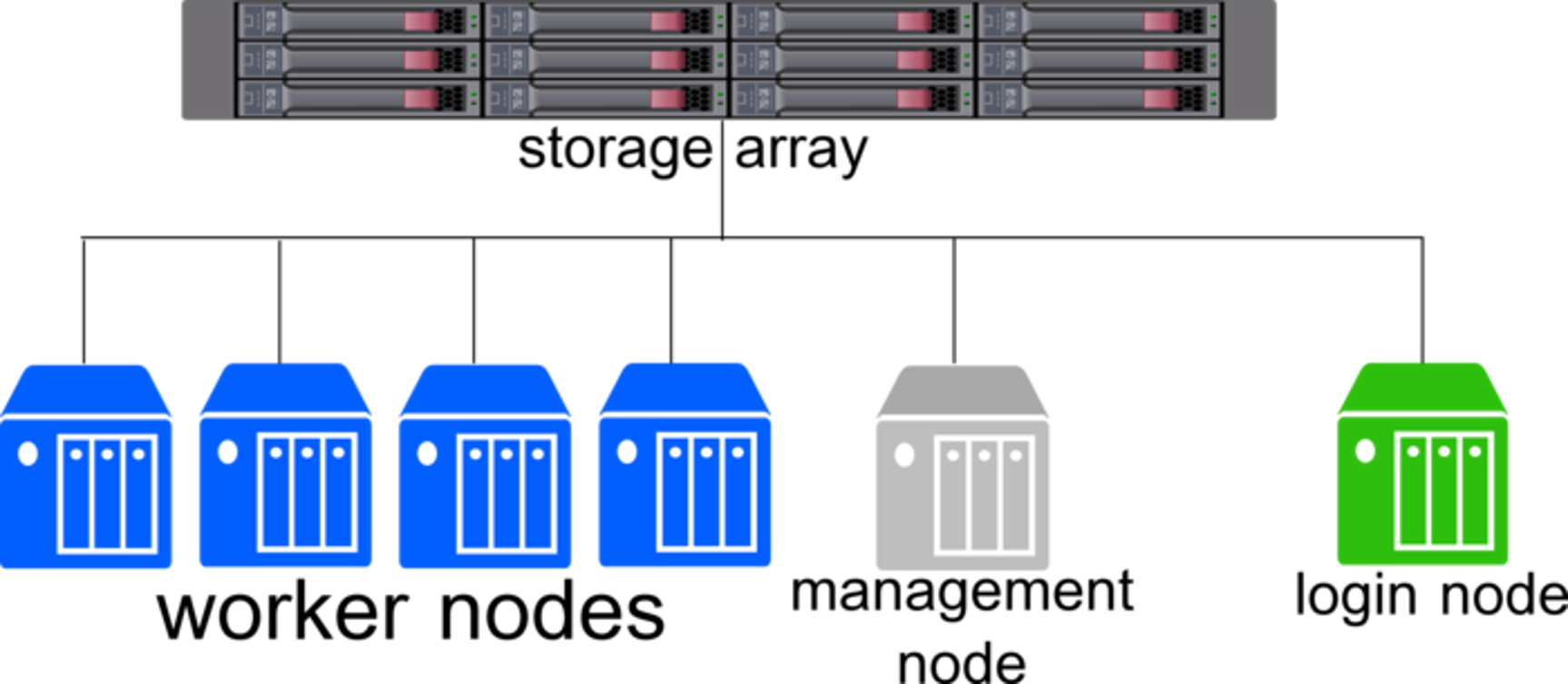
Because the Login Node is shared by everyone logging into the cluster, it is important to make sure any workflows or applications are ran as "jobs". Jobs are the unit of work on a cluster and contain all the needed components for a task to be ran on a worker node.
- The resource request - how many CPU cores, memory, GPUs, and time do you need?
- Metadata - What is the name of the job and where does the error and output go?
- Software - What software are you loading
- Workflow - What steps are needed to run your data analysis from start to finish.
Most workflows should be submitted from the command line using Slurm. HCC has some examples of submit scripts for different pieces of software available below:
More details on submitting jobs: Submitting Jobs
Loading and Using Software¶
Research computing often involves many pieces of software and different versions. Swan has over 2,000 packages pre-installed.
To help manage these packages, Swan uses a software called module to load and unload different packages.
A full list of pre-installed software is available here: Pre-Installed Software
More details on using software: Using Software
Creating custom Python, R, and Julia environments¶
Sometimes the pre-installed Python and R modules do not have the needed libraries. To do install libraries, you can use conda environments. These are environments that allow libraries to remain separate and be loaded in and out as needed.
Details on conda: Using Conda
DO NOT USE pip!
pip will cause issues with some applications on Swan. If you need to use pip, first create a conda environment and activate it first.
module purge is needed! Without it, Swan may attempt to use
# Purge all loaded modules
module purge
# Load conda
module load anaconda
# Activate conda environment
conda activate your_environment_name
# Install pip packages
pip install package_name
Integrating your custom environment into Jupyter¶
Fully custom environments and containers¶
Using JupyterLab or R Studio¶
JupyterLab is available for use within Open OnDemand as an Interactive App.
R Studio has an extra step
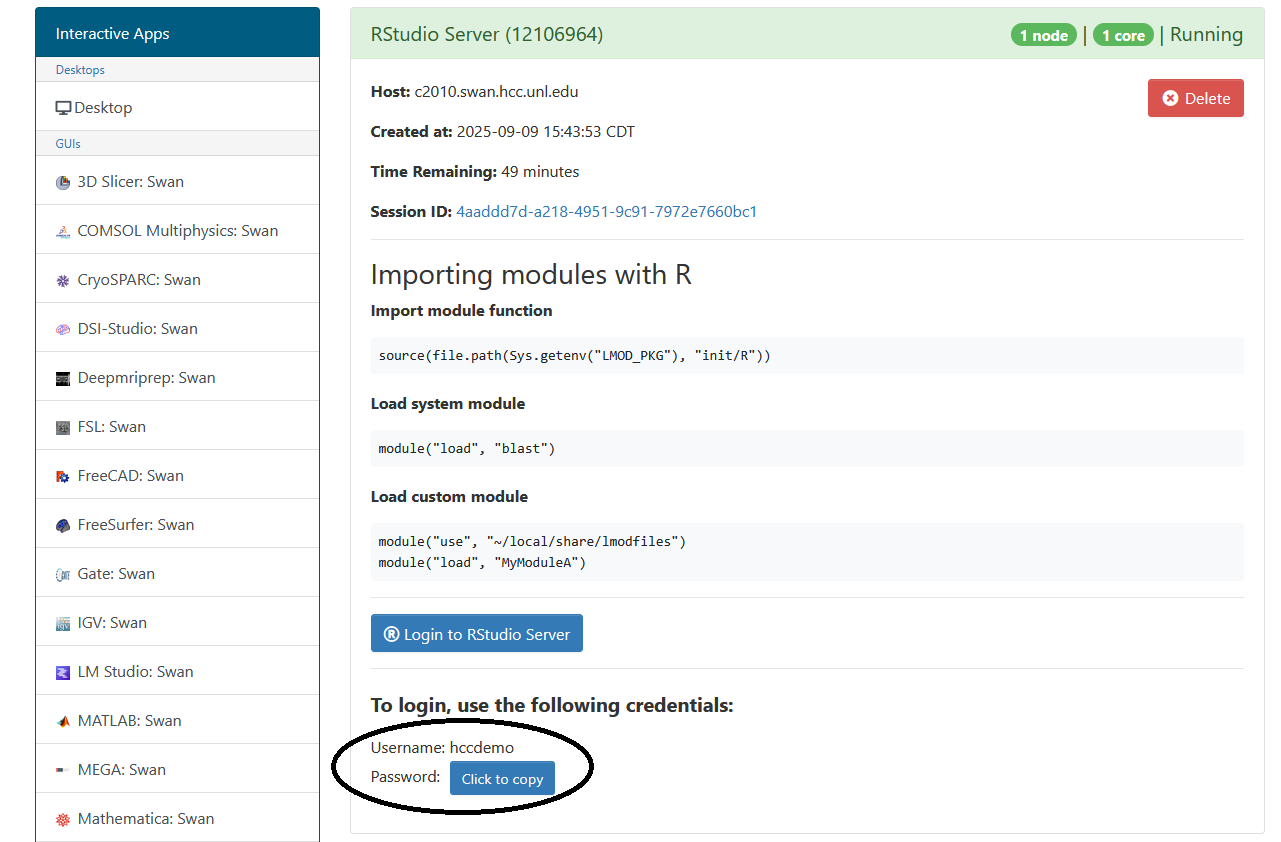
R Studio has an additional step to sign in. The application requires a different password than the one for your HCC account.
After creating the server, you will need to take note of the username and password below the "Login to RStudio Server" button. These will be your credentials for accessing the RStudio Server.
Configuration Details¶
JupyterLab and other Open OnDemand apps have a few configuration options.
| Option | Details |
|---|---|
| Version | Which version of the software you would like to use. Can be left alone. |
| Working Directory | Set it to /work/unlbayerhack/YOUR_HCC_USERNAME/ |
| Number of Cores | How many CPU cores you want. This can be a maximum of 16. |
| Running time in Hours | The time limit for the session. Can be a maximum of 8 hours. |
| Requested RAM in GBs | How much memory you want to use. Maximum of 62GB in a session. |
| Partition selection | Which set of hardware do you want to use. Set to batch or jupyter |
| Reservation | The name of a dedicated set of hardware for the hackathon. The name will be shared during the event and added here. |
| GRES | Used to request GPUs. More details on how to request a GPU is available here. |
| Job Constraints | Used to request a specfic type of GPU. More details on how to request a GPU is available here |
| If you want notifications via email of when your session starts. You will also need to click the check box below this field. |
Requesting GPUs
To request a GPU, you will need to at minimum add gpu:1 to GRES and change the Partition selection to gpu. If you want a specific type of GPU, please add the appropriate constraint to Job Constraints.
Reservations
The reservation for the hackathon will only be available from October 24th to October 26th. After October 26th, the reservation name will need removed from any scripts or session configurations.
Viewing Results and Transfering Data¶
There are multiple storage locations on Swan to store data.


- Work – High-performance global scratch (100 TiB/group, 6-month automatic purge)
- NRDStor – General-purpose (50 TiB/group, no purge)
- Home – Swan user home directories (20 GiB/user, no purge)
- Local /scratch – Node-local flash storage for running jobs
Some bioinformatics tools will require the use of /scratch. These will be highlighted when the software is loaded. As example of using /scratch is available as a part of the blast job example.
There are also multiple methods to transfer data to and from your computer. For the hackathon using a graphical application such as WinSCP or Cyberduck may be beneficial.
NRDStor
NRDStor is able to be accessed directly from your laptop in Finder (MacOS) or File Explorer (Windows) as if it was an external hard drive.
If you wish to mount NRDStor, you will need to complete the full process of gaining SMB access to NRDStor in advance of Saturday afternoon. You will need to complete the following:
Both steps will need to be completed before Saturday afternoon to gain access as HCC staff will need to apply the correct permissions.
Additional Materials¶
HCC has extra training materials available to review: