Running SAS on HCC
This quick start demonstrates how to use a SAS program on HCC resources through the command line and HCC OnDemand. The sample code and submit scripts can be downloaded from HCC's job-examples git repository.
Info
SAS is licensed software, and SAS on HCC resources is supported under UNL system-wide SAS license.
If you are part of UNL and would like to use SAS on HCC during the subscription period, please contact hcc-support@unl.edu to request access.
SAS on HCC Clusters¶
SAS applications can be run on HCC clusters similar to other jobs.
Connect to an HCC cluster and make a subdirectory
called sas_demo under your $WORK directory.
In the subdirectory sas_demo, save the SAS code. Here, we include a single demo
programs, t_test.sas, to perform a t-test analysis on a small data set.
Creating a Submit Script¶
Create a submit script to request one core (default) and 10 minutes run time on the supercomputer. The name of the main program is entered at the last line.
Submit the Job¶
The job can be submitted through the command sbatch. The job status
can be monitored by entering squeue with the -u option.
$ sbatch sas.submit
$ squeue -u <username>
Replace <username> with your HCC username.
Sample Output¶
The results of the t-test are computed and printed to the .lst file
SAS on HCC OnDemand¶
On HCC OnDemand, SAS can be accessed via Jupyter Lab Notebook and dedicated SAS Interactive GUI Application, visible for researchers with valid SAS license.
SAS on Jupyter Lab¶
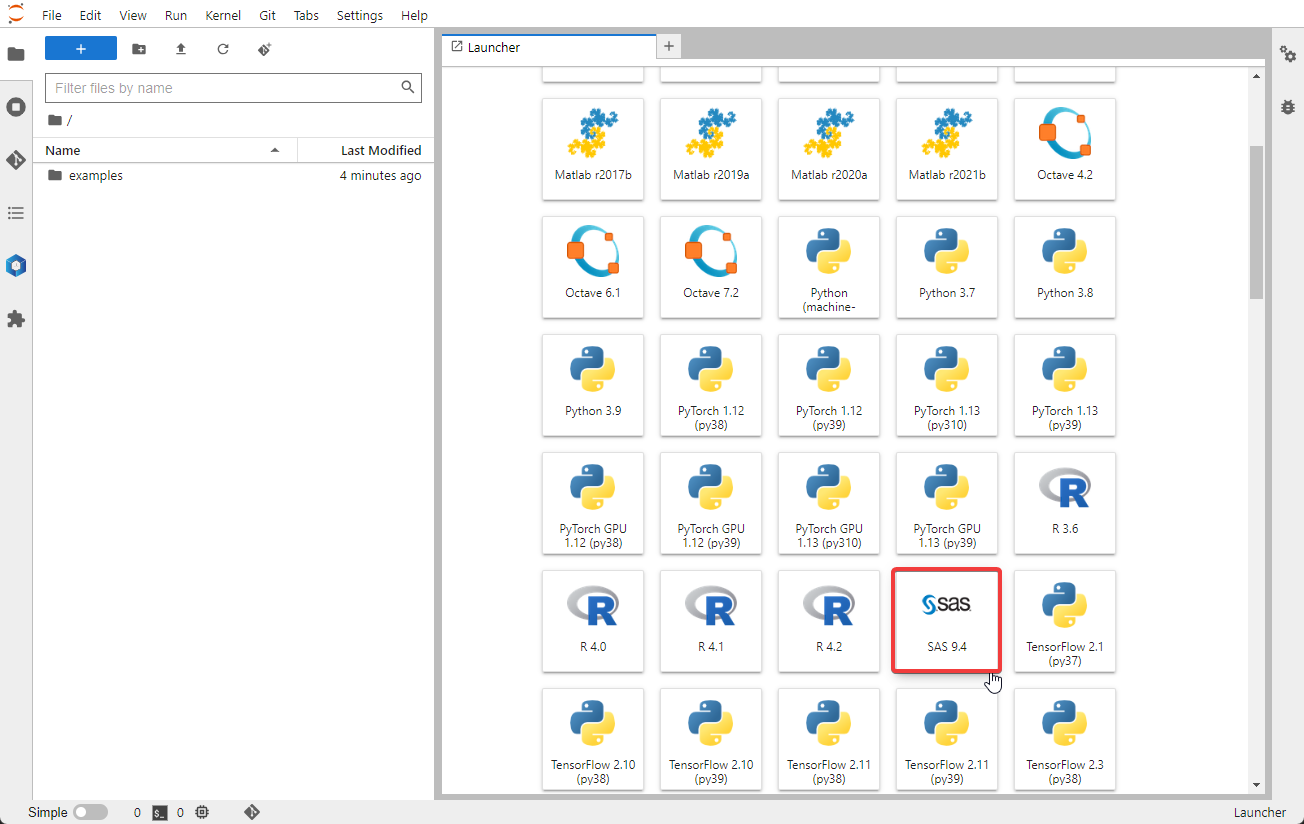
Launch a Jupyter Lab Notebook session. After the Jupyter Lab Notebook session starts, select SAS from the New dropdown box.
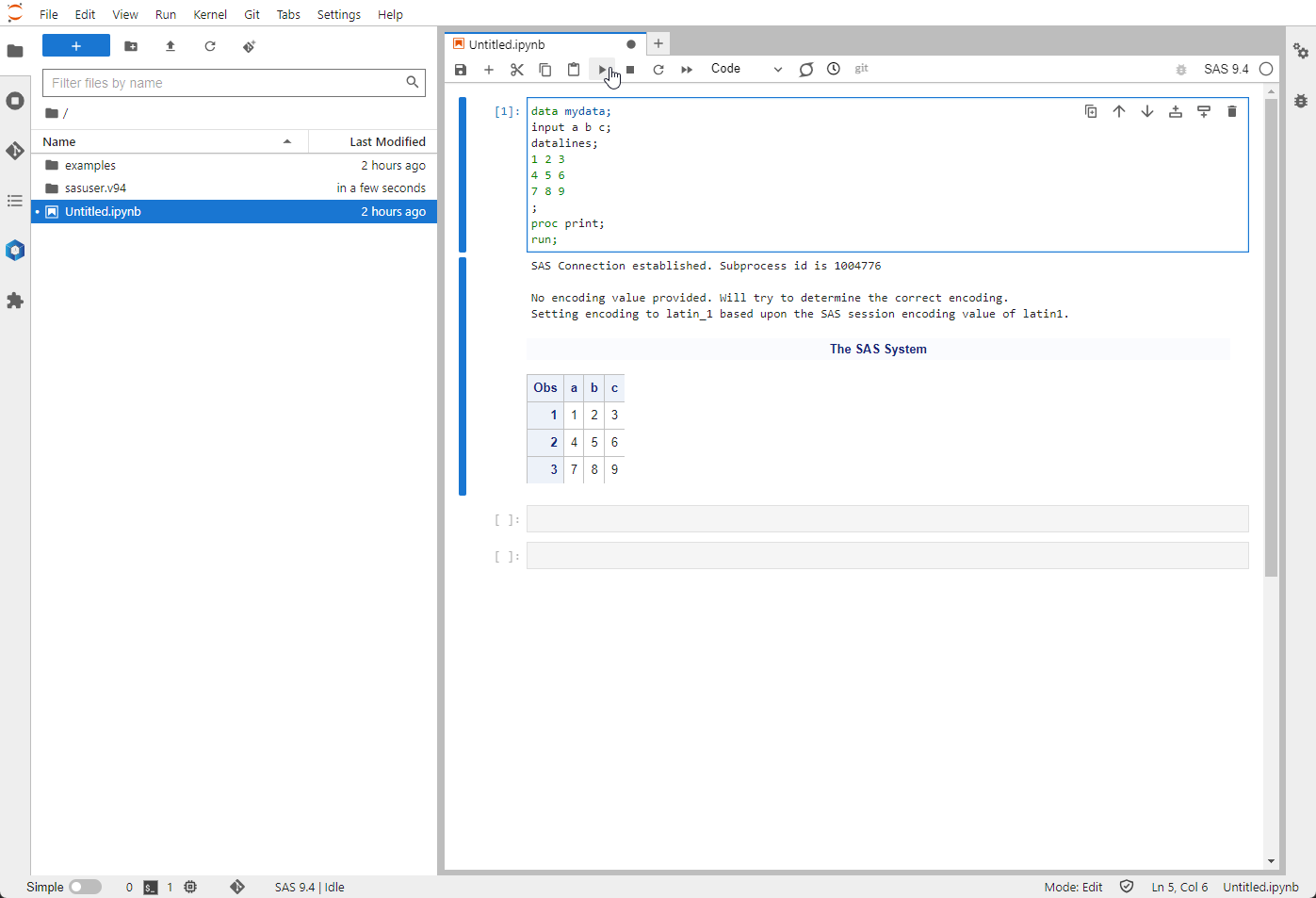
 Here, you can run code in the Notebook's cells. The SAS code is then executed when you click on the "play" icon or press the
Here, you can run code in the Notebook's cells. The SAS code is then executed when you click on the "play" icon or press the shift and enter keys simultaneously.
SAS Interactive GUI Application¶
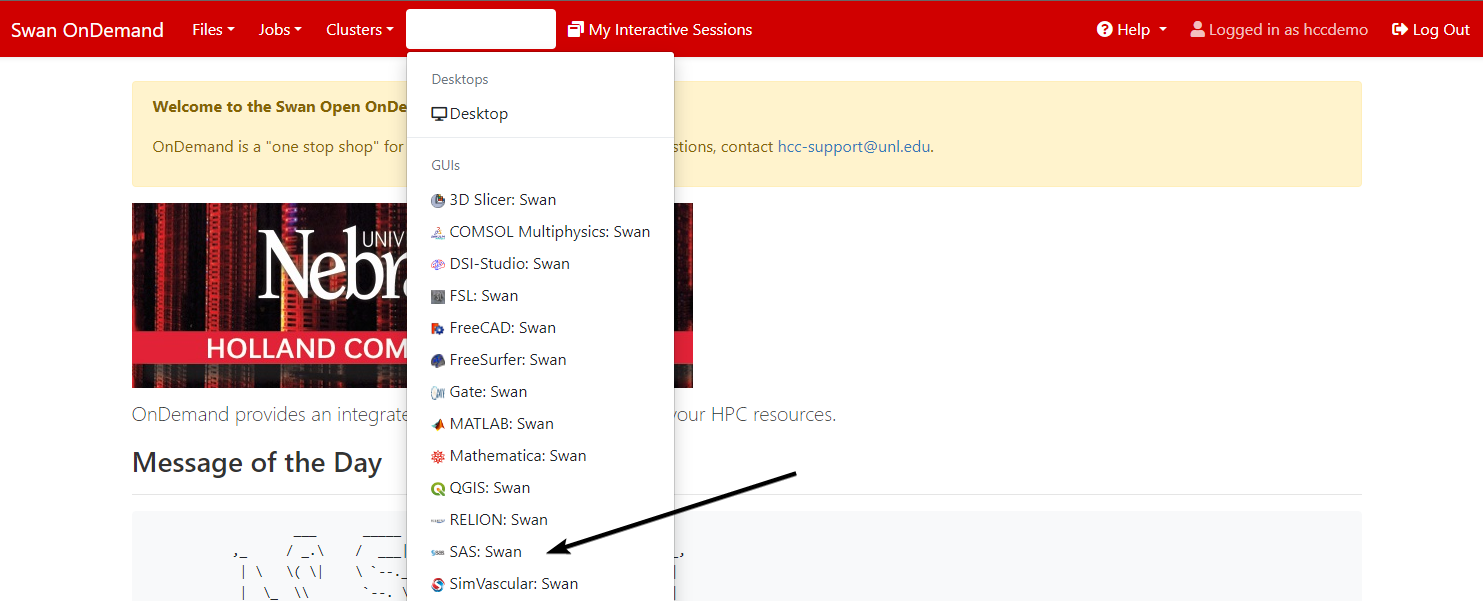
Launch the Interactive App by selecting SAS from the Interactive Apps drop-down menu at the top of the OnDemand Dashboard page and filling in the parameters needed for your job.
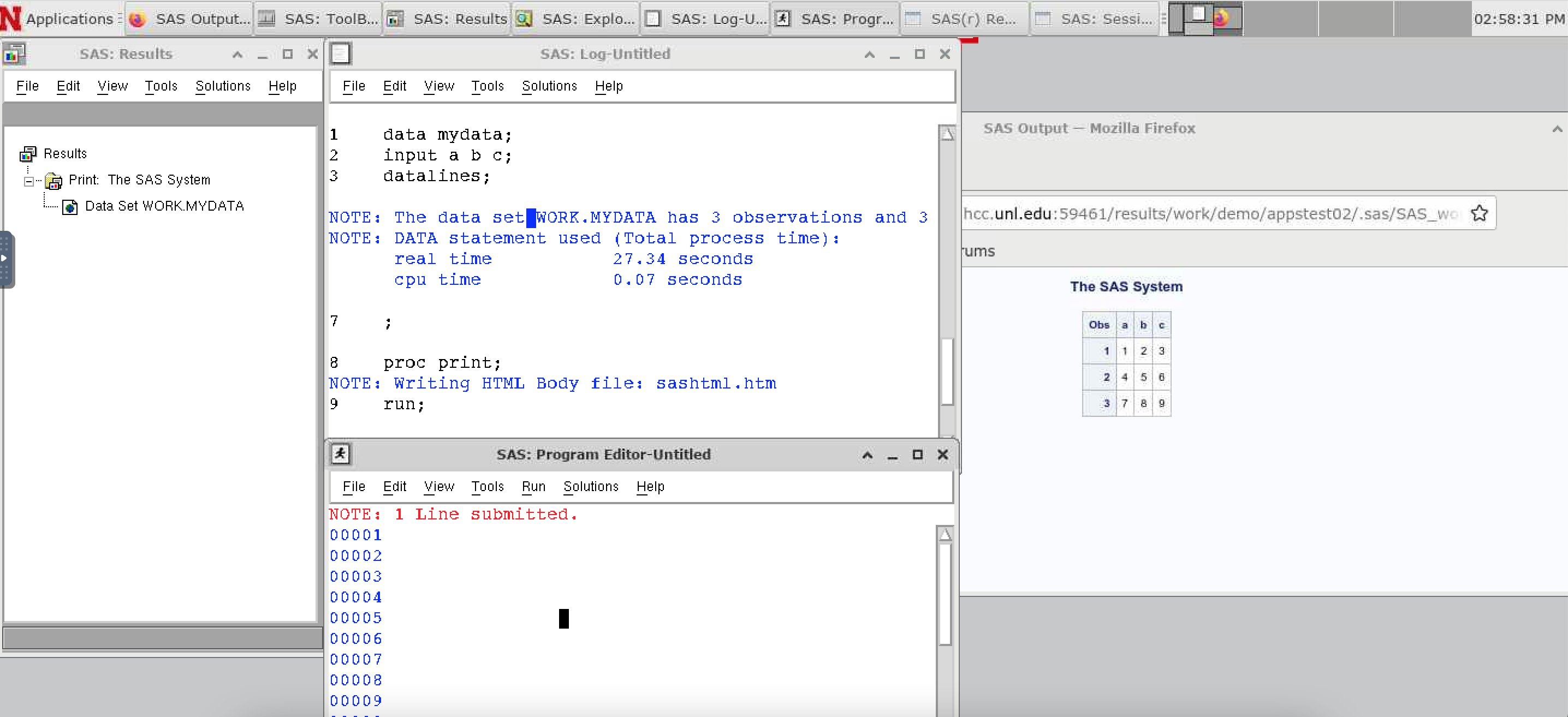
 SAS scripts can then be run from the Editor, and the results will be shown in a browser.
SAS scripts can then be run from the Editor, and the results will be shown in a browser.